Genetics
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Formats
Representation types
Update frequencies
status
Scale
-
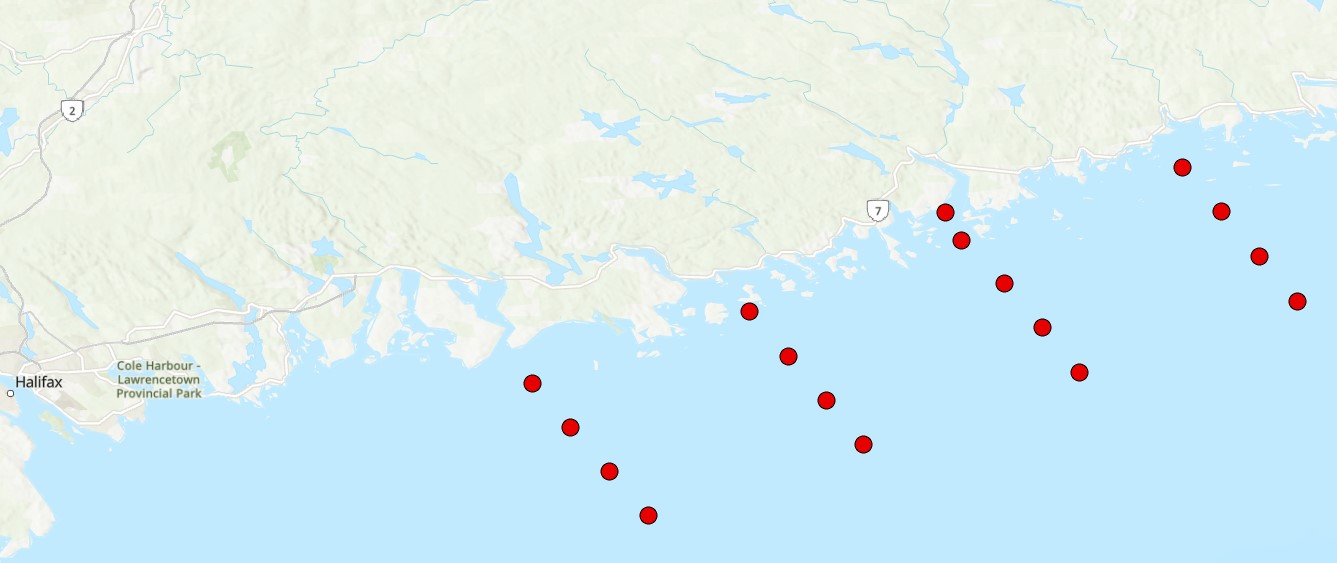
The Eastern Shore Islands Area of Interest (AOI) is a large coastal AOI in the Maritimes Region spanning 2089 km2 and extending along 100 km of coastline. While much of the data that led to the area's designation as an AOI was collected in the nearshore coastal areas (i.e., seagrass and kelp distributions, coastal bird nesting sites, herring spawning areas), there is less information available on the fish and invertebrate community structure in offshore areas of the AOI. Current offshore sampling effort is primarily focused in the Summer multi-species RV survey which covers the Scotian Shelf but does not sample within the depth range covered by the AOI (~ <100m depth). Environmental DNA (eDNA) offers a promising approach for biodiversity monitoring that has been gaining increased attention in the marine realm. Through sampling relatively small volumes of water (~1-5 L), eDNA can capture of millions of fragments of eDNA on filters that can then be sequenced to identify the species composition of an area. eDNA metabarcoding is based on the foundations of DNA barcoding, where species are identified solely by their unique DNA sequences. eDNA metabarcoding relies on a comprehensive reference library of gene sequences from known species to rapidly identify which species are captured in every water sample. This method is relatively simple, efficient (with the ability to characterize both fish and invertebrate diversity), and non-invasive, meaning there is no disruption of sensitive benthic habitats or need to dissect tissue samples from captured fish and invertebrates. eDNA has been shown to be comparable to other biodiversity censusing techniques and has the potential to rapidly conduct biodiversity surveys over a relatively large area, such as the Eastern Shore Islands AOI. Our eDNA sampling in the Eastern Shore Islands targets fish and invertebrates using multiple genetic markers (e.g., 12S and COI) to obtain baseline information on these communities across transects spanning from inshore to offshore in the AOI. Over time, we will investigate changes in species richness and community composition using annual eDNA surveys as an ongoing monitoring tool for this coastal region. Cite this data as: Jeffery, N.W. Environmental DNA Monitoring in the Eastern Shore Islands Area of Interest - Offshore. Published: April 2026. Coastal Ecosystems Science Division, Maritimes Region, Fisheries and Oceans Canada, Dartmouth NS.
-
We evaluated an autonomous environmental DNA sampler produced by Dartmouth Ocean Technologies Inc (Dartmouth, Canada) compared to time-at-sample filtration in the laboratory to determine the performance of moored samplers for monitoring in the marine world. We deployed three autonomous samplers from DOT in the Bedford Basin (Canada) over a nine-week period in summer/fall 2023. The samplers filtered seawater in situ at programmed interviews over this time period, and we collected contemporaneous samples with a standard vacuum pump during each sampling period. Both eDNA sample types captured similar fish diversity, including typical diversity for the Northwest Atlantic. The invertebrate community detected using the COI marker was different between each sample type, likely due to differences in filter pore size. We found biofouling on the moored samplers was minimal over the study period, even in a high-traffic area such as the Bedford Basin, likely due to the relatively short experimental period, and copper screening covering in the inlet and outlet valves of the instruments. Overall, our results show promise to deploy autonomous eDNA samplers in marine conservation areas to contribute to monitoring in the temperate ocean, but further testing over longer periods of time is needed to determine if DNA remains well-preserved in the autonomous samplers at ambient ocean temperatures. Cite this data as: Jeffery, N.W., Van Wyngaarden, M., and Stanley, R.R.E. Evaluating an Autonomous eDNA Sampler for Marine Environmental Monitoring: Short- and Long-Term Applications. Published: December 2024. Coastal Ecosystems Science Division, Maritimes Region, Fisheries and Oceans Canada, Dartmouth NS.
-
Appendix 17.3. Phylogeographic and population genetics studies of selected Arctic species.
-
To assess the current distribution of kelp beds and other macroalgae in Nova Scotia and Southwest New Brunswick, subtidal drop camera surveys were performed from 2022-2023 at 140 sites. For each site, a GoPro HERO 10 camera was towed along a deep (7-12m) and shallow (3-5m) depth contour until 20 images were acquired per depth. Where possible, species were identified from photos, with particular care given to kelps (defined here as orders Laminariales and Tilopteridales) and fucoids (order Fucales). Crust-forming algae was not counted. Percent cover was calculated using a 10x10 point grid overlaid on each image and recording the dominant cover type at each point. The depth (in meters) of each photo after correcting for tide height ranged from ~0.5m to ~12m. Depths were corrected to chart datum (lowest astronomical tide) using tide predictions from the nearest tide station, taken from tides.gc.ca. Sampling was performed between July and October. Cite this data as: Krumhansl K, Brooks C, Lowen B, DiBacco C, (2025). Camera Surveys of the Subtidal Flora of Nova Scotia and Southwest New Brunswick 2022-2023. Version 1.7. Fisheries and Oceans Canada. Samplingevent dataset. https://ipt.iobis.org/obiscanada/resource?r=camera_surveys_of_the_subtidal_flora_of_nova_scotia_2022-2023&v=1.7 For additional information please see: Krumhansl K.A., Brooks C.M., Lowen B., O’Brien J., Wong M., DiBacco C. Loss, resilience and recovery of kelp forests in a region of rapid ocean warming. Annals of Botany 2024 Mar 8; 133(1):73-92. Brooks C.M., Krumhansl K.A. 2023. First record of the Asian Antithamnion sparsum Tokida, 1932 (Ceramiales, Rhodophyta) in Nova Scotia, Canada. BioInvasions Records 12(3):745-725.
-
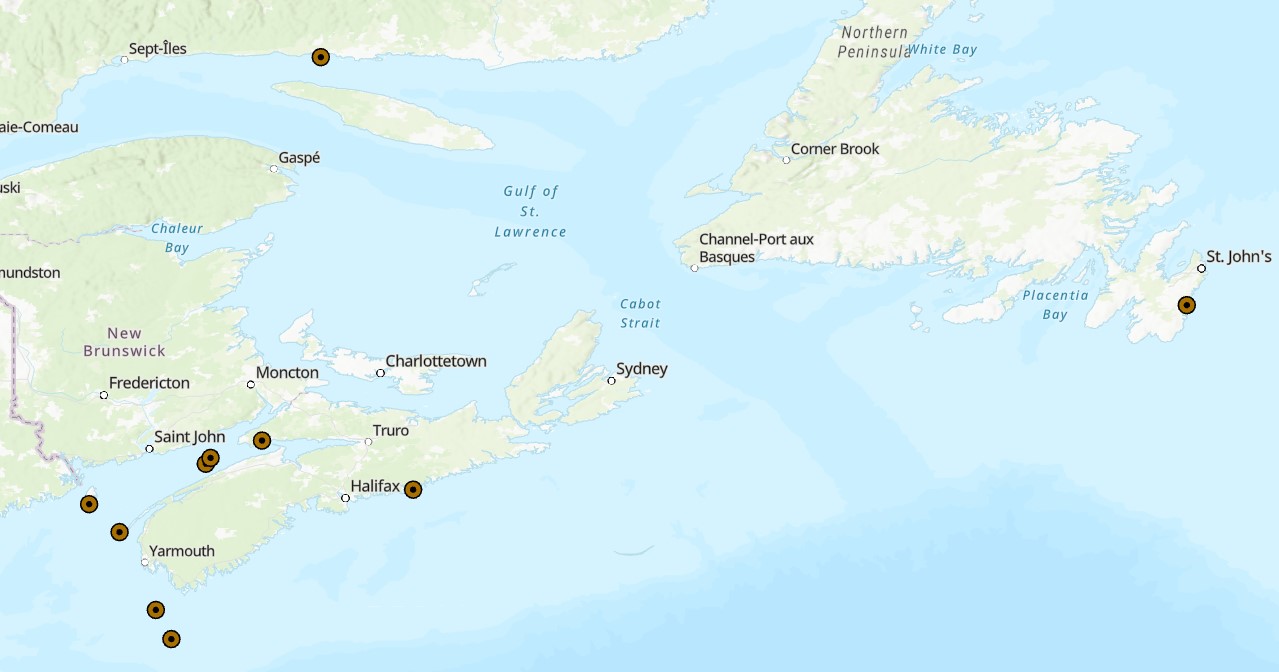
Effective conservation planning relies on understanding population connectivity which can be informed by genomic data. This is particularly important for sessile species like the horse mussel (Modiolus modiolus), a key habitat-forming species and conservation priority in Atlantic Canada), yet little genomic information is available to describe horse mussel connectivity patterns. We used more than 8000 restriction-site associated DNA sequencing-derived single nucleotide polymorphisms and a panel of 8 microsatellites to examine genomic connectivity among horse mussel populations in the Bay of Fundy, along the Scotian Shelf, and in the broader northwestern Atlantic extending to Newfoundland. Despite phenotypic differences between sampling locations, we found an overall lack of genetic diversity and population structure in horse mussels in the Northwest Atlantic Ocean. All sampled locations had low heterozygosity, very low FST, elevated inbreeding coefficients, and deviated from Hardy-Weinberg Equilibrium, highlighting generally low genetic diversity across all metrics. Principal components analysis, Admixture analysis, pairwise FST calculations, and analysis of outlier loci (potentially under selection) all showed no independent genomic clusters within the data, and an analysis of molecular variance showed that less than 1% of the variation within the SNP dataset was found between sampling locations. Our results suggest that connectivity is high among horse mussel populations in the Northwest Atlantic, and coupled with large effective population sizes, this has resulted in minimal genomic divergence across the region. These results can inform conservation design considerations in the Bay of Fundy and support further integration into the broader regional conservation network. Cite this data as: Van Wyngaarden, Mallory et al. (2024). Widespread genetic similarity between Northwest Atlantic populations of the horse mussel, Modiolus modiolus. Published: May 2025. Coastal Ecosystem Science Division, Maritimes Region, Fisheries and Oceans Canada, Dartmouth, NS.
-
A global decline in seagrass populations has led to renewed calls for their conservation as important providers of biogenic and foraging habitat, shoreline stabilization, and carbon storage. Eelgrass (Zostera marina) occupies the largest geographic range among seagrass species spanning a commensurately broad spectrum of environmental conditions. In Canada, eelgrass is managed as a single phylogroup despite occurring across three oceans and a range of ocean temperatures and salinity gradients. Previous research has focused on applying relatively few markers to reveal population structure of eelgrass, whereas a whole genome approach is warranted to investigate cryptic structure among populations inhabiting different ocean basins and localized environmental conditions. We used a pooled whole-genome re-sequencing approach to characterize population structure, gene flow, and environmental associations of 23 eelgrass populations ranging from the Northeast United States, to Atlantic, subarctic, and Pacific Canada. We identified over 500,000 SNPs, which when mapped to a chromosome-level genome assembly revealed six broad clades of eelgrass across the study area, with pairwise FST ranging from 0 among neighbouring populations to 0.54 between Pacific and Atlantic coasts. Genetic diversity was highest in the Pacific and lowest in the subarctic, consistent with colonization of the Arctic and Atlantic oceans from the Pacific less than 300 kya. Using redundancy analyses and two climate change projection scenarios, we found that subarctic populations are predicted to be more vulnerable to climate change through genomic offset predictions. Conservation planning in Canada should thus ensure that representative populations from each identified clade are included within a national network so that latent genetic diversity is protected, and gene flow is maintained. Northern populations, in particular, may require additional mitigation measures given their potential susceptibility to a rapidly changing climate. Cite this data as: Jeffery, Nicholas et al. (2024). Data from: Variation in genomic vulnerability to climate change across temperate populations of eelgrass (Zostera marina) [Dataset]. https://doi.org/10.5061/dryad.xpnvx0kp2
-

An exploratory project on the taxonomic and genetic diversity of decapods in three ocean subregions (Northeast Pacific, Canadian Arctic, and Northwest Atlantic), which were sampled in 2022, was undertaken by the Arctic Working Group under the Canada-U.S. Fisheries and Climate Collaboration between Fisheries and Oceans Canada (DFO) and the National Marine Fisheries Service (NMFS) of the National Oceanic and Atmospheric Administration (NOAA). This collaboration framework aims to pool Canadian and U.S. data to explore the impacts of broad-scale climate change on marine biodiversity. In early summer 2022, a sampling protocol with the selection of targeted decapods was provided to DFO and NOAA collaborators. Targeted genera were collected from a total of 10 research programs across three ocean subregions and four marine regions. The Northeast Pacific samples were collected in the Bering Sea during the Northern Bering Sea Ecosystem and Surface Trawl Survey, and the Eastern and Northern Bering Sea Continental Shelf Bottom Trawl Survey of Groundfish and Invertebrate Fauna onboard the F/V Northwest Explorer, F/V Alaska Knight and F/V Vesteraalen. In the Western Canadian Arctic (mainly from Beaufort Sea and Amundsen Gulf), specimens were collected during DFO’s Canadian Beaufort Sea – Marine Ecosystem Assessment (CBS-MEA) survey onboard the F/V Frosti. In Eastern Canadian Arctic (mainly from Baffin Bay and Davis Strait), specimens were collected during DFO’s Knowledge and Ecosystem-Based Approach in Baffin Bay (KEBABB) survey onboard the CCGS Amundsen and DFO’s North Atlantic Fisheries Organization (NAFO) Subarea 0B survey onboard the R/V Tarajoq. In the Estuary and Gulf of St. Lawrence (EGSL), specimens were collected from coastal surveys (scallops, sea cucumber, snow crab, and whelk surveys) onboard the CCGS Leim and offshore during the Ecosystemic Survey onboard the CCGS Teleost. Decapods were collected from various sampling gears (benthic beam trawl, modified Atlantic Western IIA otter trawl, Bacalao trawl, shrimp trawl, Digby scallop dredge, or modified sea cucumber dredge) and identified to the lowest possible taxonomic level and photographed, when possible. All specimens were frozen at sea (n = 995). In the lab, the identifications were validated or refined with the photos and the frozen specimens. DNA was extracted for 87 specimens and a section of COI gene was amplified in order to be sequenced using Sanger method. Sequences were compared with existing data using The Basic Local Alignment Search Tool (BLAST) in the National Center for Bio-technology Information Nucleotide database (NCBI-nt, including the GenBank database) to compare scientific names, where available. The present dataset includes 391 decapod species occurrences. DNA was extracted for a subset of 87 specimens (COI gene); sequences are publicly available on BOLD data portal under project code DDAO (see supporting document "citations_references.csv" for more information). The data are presented in Darwin Core format and are separated in three files: The "Activité_décapodes_DDAO_decapods_event_en" file contains information about missions, stations and deployments, which are presented under a hierarchical activity structure. The "Occurrence_décapodes_DDAO_decapods_en" file contains the taxonomic occurrences. The "ADN_décapodes_DDAO_decapods_DNA_en" file contains the DNA derived data. For further details, please refer to the technical report available in the supporting document named "citations_references.csv". USE LIMITATION: To ensure scientific integrity and appropriate use of the data, we would encourage you to contact the data custodian.
 Arctic SDI catalogue
Arctic SDI catalogue