oceans
Type of resources
Available actions
Topics
Keywords
Contact for the resource
Provided by
Years
Formats
Representation types
Update frequencies
status
Scale
Resolution
-

The Regional Deterministic Wave Prediction System (RDWPS) produces wave forecasts out to 48 hours in the future using the third generation spectral wave forecast model WaveWatch III® (WW3). The model is forced by the 10 meters winds from the High Resolution Deterministic Prediction System (HRDPS). Over the Great Lakes, an ice forecast from the Water Cycle Prediction System of the Great Lakes (WCPS) is used by the model to attenuate or suppress wave growth in areas covered by 25% to 75% and more than 75% ice, respectively. Over the ocean, an ice forecast from the Regional Ice Ocean Prediction System (RIOPS) is used: in the Northeast Pacific, waves propagate freely for ice concentrations below 50%, above this threshold there is no propagation; in the Northwest Atlantic the same logic is used as in the Great Lakes. Forecast elements include significant wave height, peak period, partitioned parameters and others. This system includes several domains: Lake Superior, Lake Huron-Michigan, Lake Erie, Lake Ontario, Atlantic North-West and Pacific North-East.
-
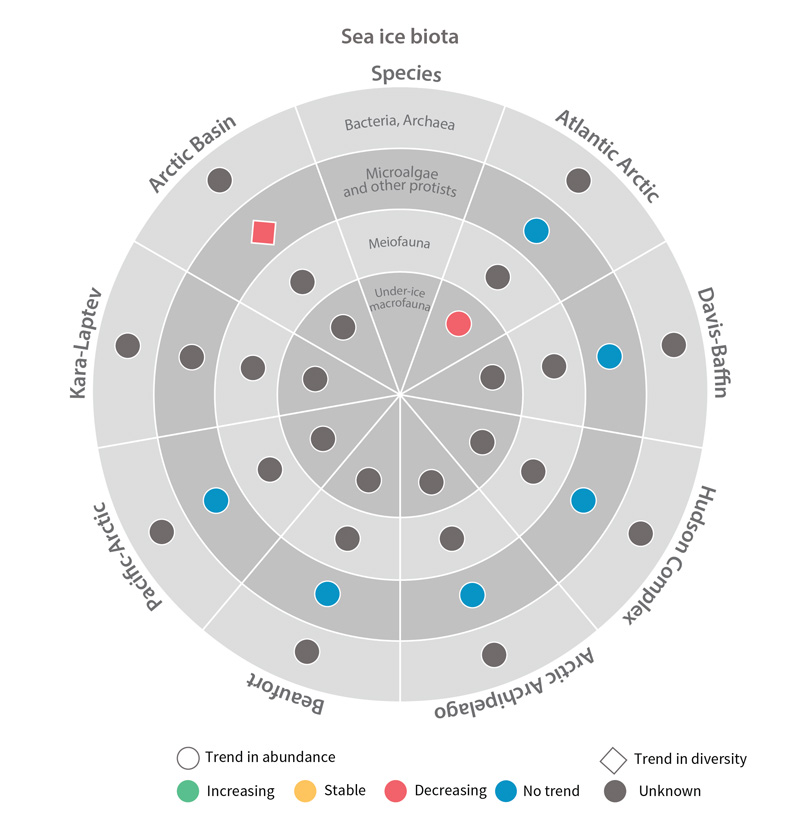
Trends in abundance or diversity of sea ice biota Focal Ecosystem Components across each Arctic Marine Area. STATE OF THE ARCTIC MARINE BIODIVERSITY REPORT - Chapter 4 - Page 177 - Figure 4.1
-
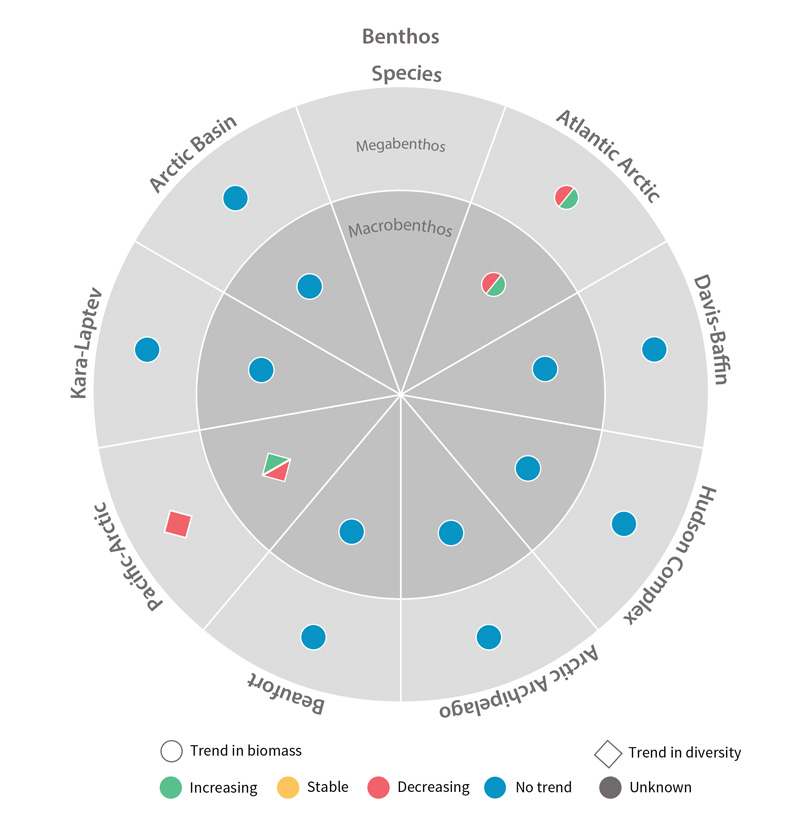
Trends in biomass or diversity of benthic Focal Ecosystem Components across each Arctic Marine Area. STATE OF THE ARCTIC MARINE BIODIVERSITY REPORT - Chapter 4 - Page 179 - Figure 4.3
-
Water body silicate - Monthly Climatology for the European Seas for the period 1960-2020 on the domain from longitude -45.0 to 70.0 degrees East and latitude 24.0 to 83.0 degrees North. Data Sources: observational data from SeaDataNet/EMODnet Chemistry Data Network. Description of DIVA analysis: The computation was done with the DIVAnd (Data-Interpolating Variational Analysis in n dimensions), version 2.7.2, using GEBCO 30sec topography for the spatial connectivity of water masses. Horizontal correlation length and vertical correlation length vary spatially depending on the topography and domain. Depth range: 0.0, 5.0, 10.0, 15.0, 20.0, 25.0, 30.0, 35.0, 40.0, 45.0, 50.0, 55.0, 60.0, 65.0, 70.0, 75.0, 80.0, 85.0, 90.0, 95.0, 100.0, 125.0, 150.0, 175.0, 200.0, 225.0, 250.0, 275.0, 300.0, 325.0, 350.0, 375.0, 400.0, 425.0, 450.0, 475.0, 500.0, 550.0, 600.0, 650.0, 700.0, 750.0, 800.0, 850.0, 900.0, 950.0, 1000.0, 1050.0, 1100.0, 1150.0, 1200.0, 1250.0, 1300.0, 1350.0, 1400.0, 1450.0, 1500.0, 1550.0, 1600.0, 1650.0, 1700.0, 1750.0, 1800.0, 1850.0, 1900.0, 1950.0, 2000.0, 2100.0, 2200.0, 2300.0, 2400.0, 2500.0, 2600.0, 2700.0, 2800.0, 2900.0, 3000.0, 3100.0, 3200.0, 3300.0, 3400.0, 3500.0, 3600.0, 3700.0, 3800.0, 3900.0, 4000.0, 4100.0, 4200.0, 4300.0, 4400.0, 4500.0, 4600.0, 4700.0, 4800.0, 4900.0, 5000.0, 5100.0, 5200.0, 5300.0, 5400.0, 5500.0 m. Units: umol/l. The horizontal resolution of the produced DIVAnd analysis is 0.25 degrees.
-
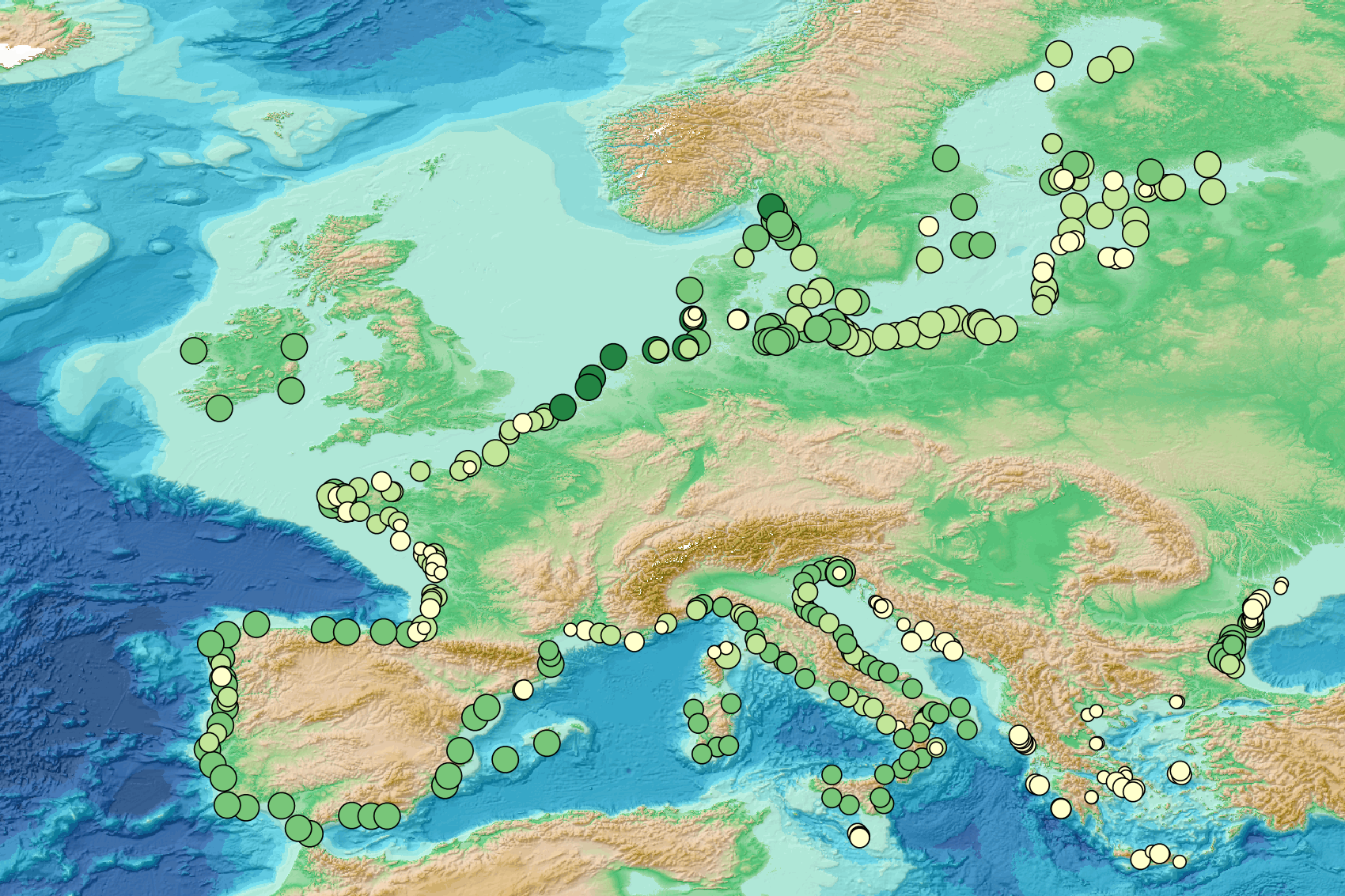
This visualization product displays the number of Marine Strategy Framework Directive (MSFD) monitoring surveys and the associated temporal coverage per beach. EMODnet Chemistry included the collection of marine litter in its 3rd phase. Since the beginning of 2018, data of beach litter have been gathered and processed in the EMODnet Chemistry Marine Litter Database (MLDB). The harmonization of all the data has been the most challenging task considering the heterogeneity of the data sources, sampling protocols and reference lists used on a European scale. Preliminary processing were necessary to harmonize all the data: - Exclusion of OSPAR 1000 protocol: in order to follow the approach of OSPAR that it is not including these data anymore in the monitoring; - Selection of MSFD surveys only (exclusion of other monitoring, cleaning and research operations); - Exclusion of beaches without coordinates. More information is available in the attached documents. Warning: the absence of data on the map doesn't necessarily mean that they don't exist, but that no information has been entered in the Marine Litter Database for this area.
-
In 2012 and 2013, Fisheries and Oceans Canada surveyed the benthos in two areas closed to bottom contact fishing, the Narwhal Overwintering and Coldwater Coral Zone (now the Disko Fan Conservation Area, DFCA), and the Hatton Basin Voluntary Coral Protection Zone (now the Hatton Basin Conservation Area, HBCA). Samples were collected following protocols recommended by the Arctic Council’s Circumpolar Biodiversity Monitoring Plan for the purposes of providing baseline data for future monitoring of benthic invertebrates in this sensitive region, and for facilitating pan-Arctic comparisons of benthic communities. Five biodiversity monitoring stations were established, four in the DFCA and one in the HBCA, each of which was fully sampled according to those protocols with Van Veen grabs or box corers, drop cameras and temperature recorders attached to the gear. This report summarises the grab/core-sampled benthic fauna collected during the 2012 survey of the Conservation Areas and complements another report documenting the epibenthos from the camera transects in the DFCA. Here we report on macrofauna in the 1-cm size fraction, and on foraminiferan meiofauna. The data provided is presented in the following report (see related link) : Jacobs, K., Bouchard Marmen, M., Rincón, B., MacDonald, B., Lirette, C., Gibb, O., Treble, M., and Kenchington, E. 2022. Biodiversity Monitoring Stations for Benthic Macrofauna and Meiofauna in the Disko Fan and Hatton Basin Conservation Areas. Can. Tech. Rep. Fish. Aquat. Sci. 3487: vi + 86 p. Cite this data as: Bouchard Marmen, Marieve; Rincon, Beatriz ; MacDonald, Barry; Lirette, Camille; Gibb, Olivia; Treble, Margaret ; Jacobs, Kevin; Kenchington, Ellen (2022). Biodiversity Monitoring Stations for Benthic Macrofauna and Meiofauna in the Disko Fan and Hatton Basin Conservation Areas. Published January 2023. Ocean Ecosystems Science Division, Fisheries and Oceans Canada, Dartmouth, N.S. https://open.canada.ca/data/en/dataset/b7bcff18-698b-4d40-a7bd-13d39925cbeb
-
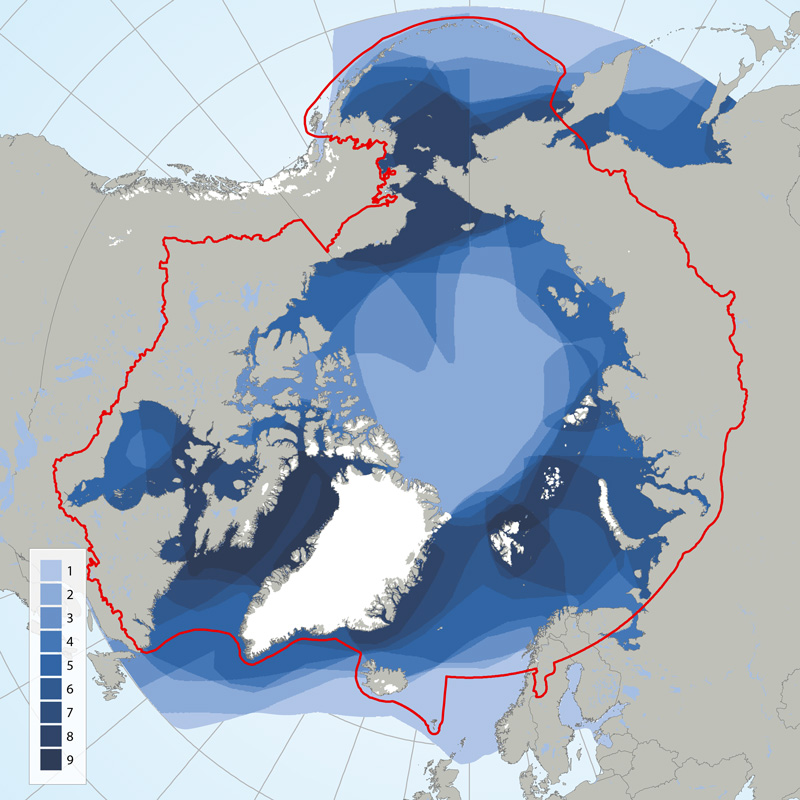
Circumpolar depiction of species richness based on the distributions of the 11 ice-associated Focal Ecosystem Components (according to the distributions reported in IUCN Red List species accounts). A maximum of nine species occur in any one geographic location. The Arctic gateways in both the Atlantic and Pacific regions have the highest species diversity. STATE OF THE ARCTIC MARINE BIODIVERSITY REPORT - <a href="https://arcticbiodiversity.is/findings/marine-mammals" target="_blank">Chapter 3</a> - Page 152 - Figure 3.6.1
-

For the background of data production and data interpretation we refer to the PAME report “Modelling Arctic oceanographic connectivity to further develop PAME’s MPA toolbox”. The uploaded data consist of two main types: 1. Connectivity matrices describing the seascape connectivity in the model domain consisting of 40893 model grid cells. The connectivity matrices describe the probability of dispersal between any two selected model grid cells. 2. GIS shape files of dispersal distance (km) from each model grid cell within the model domain.
-
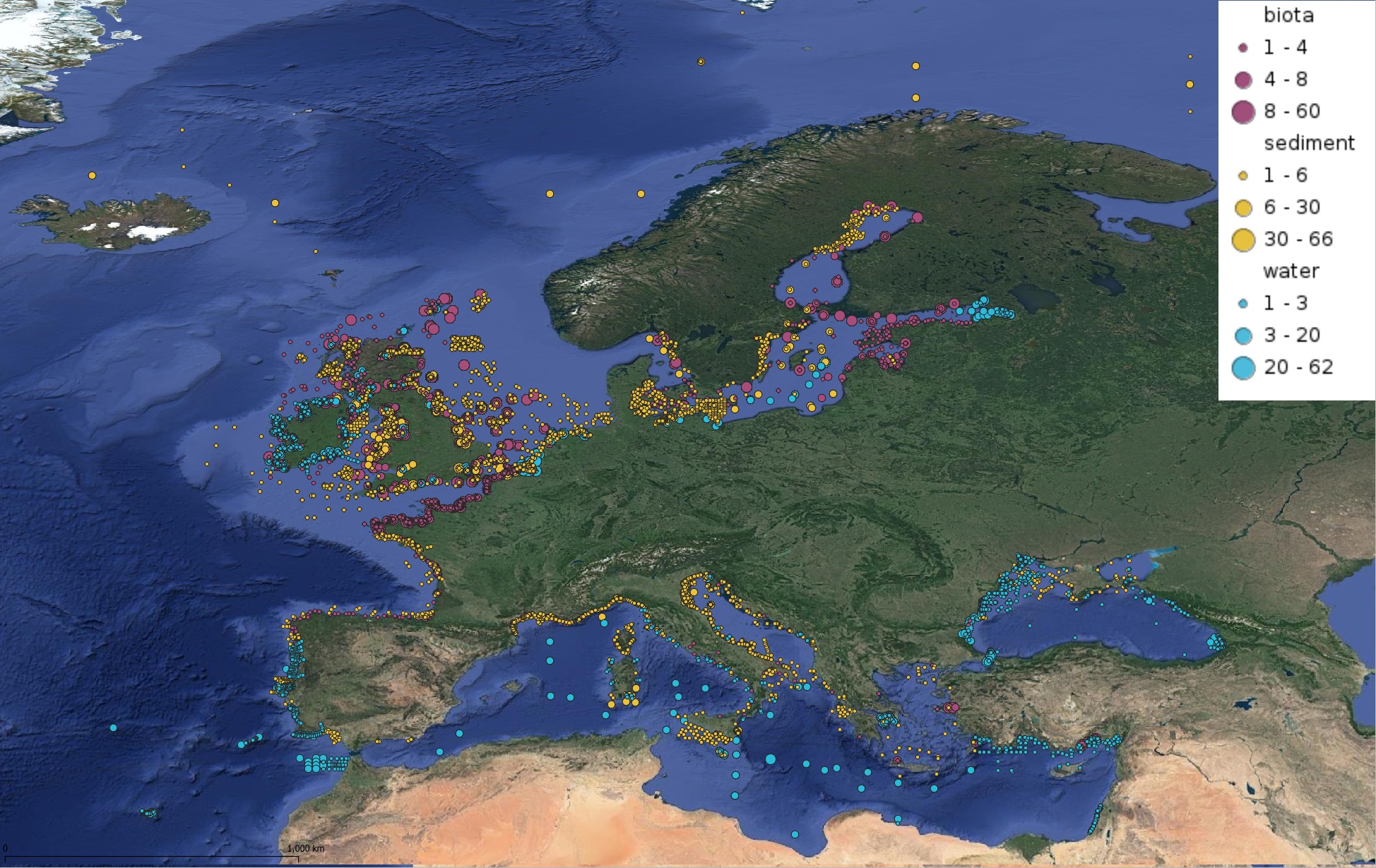
This product displays for Cadmium, positions with values counts that have been measured per matrix and are present in EMODnet regional contaminants aggregated datasets, v2022. The product displays positions for all available years.
-
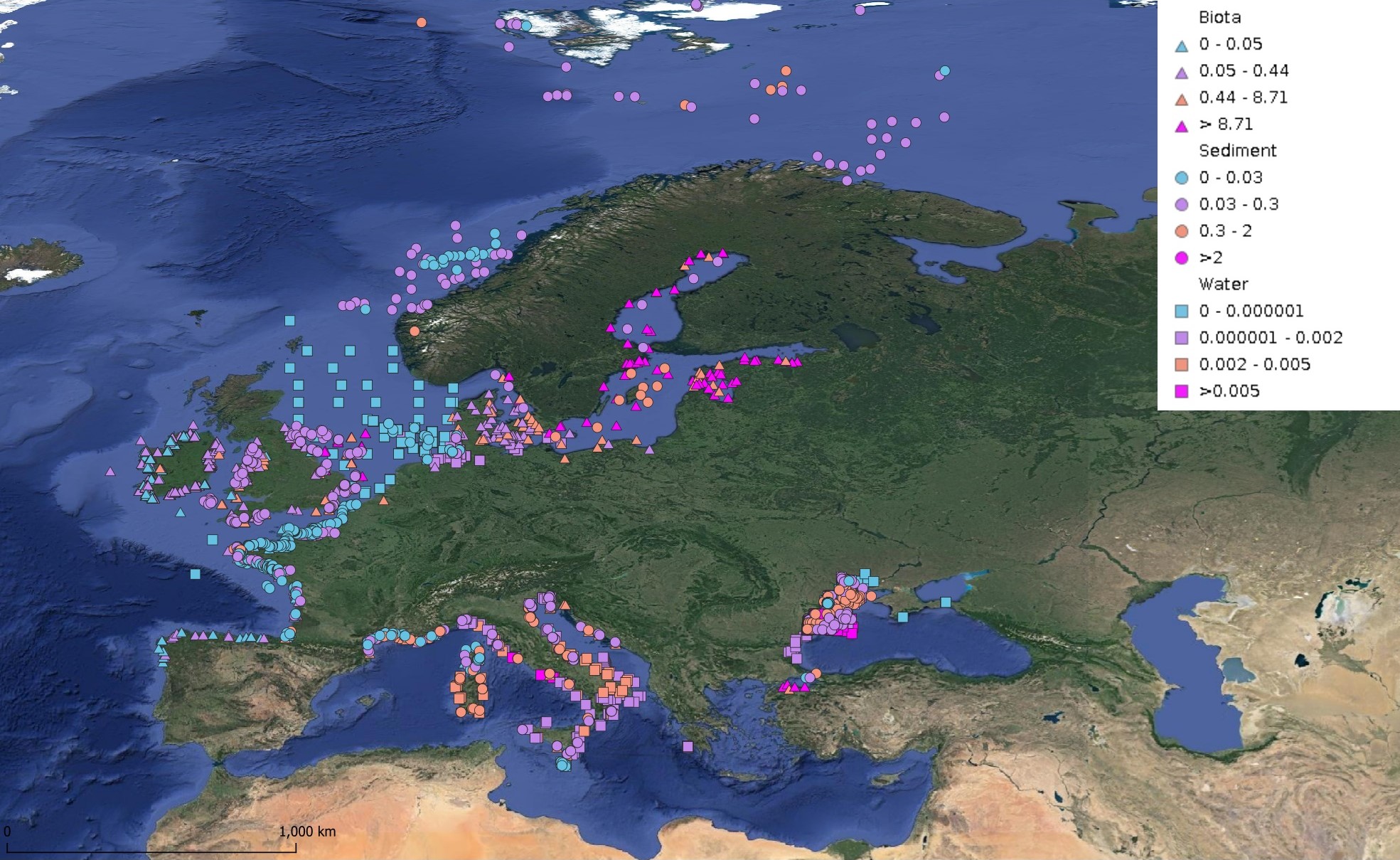
This product displays for DDT, DDE, and DDD, median values since 2012 that have been measured per matrix and are present in EMODnet regional contaminants aggregated datasets, v2024. The median values ranges are derived from the following percentiles: 0-25%, 25-75%, 75-90%, >90%. Only "good data" are used, namely data with Quality Flag=1, 2, 6, Q (SeaDataNet Quality Flag schema). For water, only surface values are used (0-15 m), for sediment and biota data at all depths are used.
 Arctic SDI catalogue
Arctic SDI catalogue